About 📝

Nikhil Nageshwar Inturi -
Sr. Data Scientist and Bioinformatician at the Center for Advanced Pain Studies | Machine Learning in Biopharma R&D and Drug Discovery
As a Senior Data Scientist with over 7 years of experience, I specialize in building high-impact, AI-driven solutions for genomics, multi-omics integration, and biomarker discovery. My expertise spans the full data science lifecycle, from developing Advanced Machine Learning and Generative AI applications to deploying scalable, containerized workflows in cloud environments.
Ever since I was a kid, I’ve been captivated by two things: how machines work, and how information flows. That curiosity led me from building my first “Hello World!” in Java to leading three amazing teams at The University of Texas at Dallas, each tackling some of the most exciting challenges in Science and AI. I’ve had the privilege of working with several distinguished researchers, including Dr. Theodore J. Price, Dr. Diana F. Tavares, and Dr. Eric Christopher Meyers at UTD, as well as collaborating with multiple labs across the globe. Before coming to the U.S., I spent 4.5 years in India's tech industry. At Infosys Ltd., I worked in consulting and development, notably for BMW and the Infosys Research Unit. There, I honed my data science skills, developing enterprise applications and algorithms, and spearheading containerization efforts that dramatically improved deployment efficiency. Later, at Aganitha, I worked as a Full-Stack Data Scientist on AAV Capsid Engineering 🧬, setting up the midstream and downstream analysis. I developed clustering models aimed at identifying optimal golden templates and predicting effective 7-mer sequences for blood-brain barrier transduction 🧠.
I have the privilege of leading teams in Bioinformatics, Generative AI, and Machine Learning for vision models. Together, we’re pushing the boundaries of what’s possible in genomics, spatial transcriptomics (Visium, Xenium), and computational biology.
- In bioinformatics, we’re making sense of massive NGS and spatial-omics datasets, accelerating biomarker discovery, and collaborating with researchers around the globe trying to decode pain.
- In Generative AI, we build Agentic-RAG solutions that help scientists with literature reviews using the latest models. I love the magic of self-attention and autoregressive architectures—it’s like teaching a computer to read, reason, and synthesize knowledge from mountains of scientific literature.
- In Machine Learning, my team builds classification (FastAI classification), object-detection (YOLO-v5, YOLO-v8), and segmentation (FastAI Segmentation, Detectron2, SAM, SAM2, Cellpose-SAM, Segment-AnyNeuron, StarDist, YOLO-v11, YOLO-v12) models that spot neurons and other cell types in high-res staining images, bringing new clarity to spatial biology and neuroscience.
I get a real thrill from translating messy, complex data—whether it’s gene sequences, images, or unstructured text—into insights that spark new discoveries. I’m hands-on with everything from Python and R to Docker, Nextflow, and cloud platforms, but what I enjoy most is mentoring others and building tools that make a difference for the explorers.
- 7+ years in Machine Learning, Generative AI, and Bioinformatics across academia and industry.
- Expert in NGS data analytics (scRNA-seq, ATAC-seq, spatial omics), multi-omics integration, and biomarker discovery.
- Advanced skills in Python, R, Nextflow, Docker, Kubernetes, and cloud (AWS, Azure).
- Developed and deployed RAG-based and multi-agent AI applications for research acceleration.
- Collaborated with researchers from McGill, UPenn, UQ-Australia, WashU, etc.
- Certified in AWS, Azure, and advanced AI/machine learning.
If you’re passionate about the intersection of AI, NLP, and life-sciences or just want to geek out about transformers and the future of data-driven discovery, let’s connect!
Work Experience 👨🏻💻
Center for Advanced Pain Studies, The University of Texas at Dallas
Feb 2023 – Present
As a Senior Data Science and Bioinformatician at Center for Advanced Pain Studies (Dr. Price, Dr. Tavares, and Dr. Meyers), I had the opportunity to work on several pipelines and interact with some amazing researchers who are exemplars in the pain field. Here is a peek into my work here as an Intern here at Center for Advance Pain Studies (CAPS):
- Built an Agentic-RAG chatbot that ingests PubMed/bioRxiv/Nature abstracts, PDFs, and internal datasets, index files from NFS, S3 and Prometheus using adaptive contextual chunking and dense hybrid search; cut evidence-gathering time by 90% for 40+ scientists at CAPS.
- Developed a multi-agent system on Azure AI Studio with 4 cooperative agents (crawl+research, reason, synthesize, publish) to surface pain-omics insights from DB tables, NFS file trees, and live web data; serves 2k queries a month with less than 200ms median latency.
- Built a neuron‑detection pipeline using image segmentation (Detectron2 | YOLOv11 | FastAI | SAM) that raised F1‑score 0.78 → 0.89 (+15%) while slashing processing time 95%.
- Deployed "Containerized Nextflow workflows" for Bulk RNA-sequencing, scRNA-seq, ATAC-seq, and spatial-omics (Visium and Xenium); scaled to 50+ sample sets on Kubernetes and accelerated biomarker discovery 60%.
- Developed ensemble models for Vagus Nerve Simulation data (XGBoost, LightGBM, LSTM), achieving 93% accuracy / R²=0.92, in predicting the rat jaw-size.
- Led 3 cross‑functional teams (15 scientists & 7 students) in image analysis, sequencing analytics, and bioinformatics; mentored 4 graduate RAs.
- Spearheaded global collaborations (McGill, UPenn, WashU and Univ. of Queensland) standardizing NGS and Spatial Transcriptomic pipelines and cutting turn‑around from weeks‑to‑days.
Aganitha Cognitive Solutions, Hyderabad, India
Jun 2022 – Dec 2022
-
I predominantly worked on *in Silico* solutions related to Gene Therapy, Drug Discovery, etc. for industry-leading clients in Health Care and Biopharma. The work here at Aganitha requires a strong understanding of molecular bioinformatics, omics, and genomics. It's quite fascinating to learn about the human body and its metabolism, and I am looking forward to learning more.
- Developed and implemented advanced algorithms for the AutoBLAST, a genome browser, doubling efficiency compared to the Basic Local Alignment Search Tool.
- Designed and executed clustering models for the AAV Capsid Engineering Project to identify the golden templates, significantly reducing the number of required in-vivo experiments by 99.96%, which led to substantial cost savings.
- Developed interactive ipython widgets and panel dashboards for the Midstream and Downstream Analysis components of AAV Capsid Engineering, enhancing user engagement and data visualization capabilities.
- Revamped Splice-AI algorithms to accurately detect and analyze novel splice junctions in humans and rodents, which resulted in a 20% increase in overall testing efficiency.
Infosys Ltd, Hyderabad, India
Sep 2018 – Jun 2022
-
During my tenure at Infosys and its research unit, Infosys ICETS, I undertook various responsibilities as mentioned below:
- Developed and implemented machine learning algorithms using a variety of packages such as scikit-learn, LGBoost, CatBoost, H2O, and keras to enhance the functionality of the Infosys Data Science and Machine Learning Platform (IDSMLP), significantly expanding the scope of the tool.
- Integrated advanced data visualization techniques into the IDSMLP tool interface by incorporating univariate, bivariate, and multivariate interactive charts using Bokeh and Plotly. Additionally, I established load balancing with an Nginx server to optimize user interactions and enhance the understanding of complex data patterns through interactive charts.
- Streamlined deployment processes by Dockerizing all applications on the ICETS-Data Science platform, which reduced deployment times by 75% and significantly increased operational efficiency.
- Developed and implemented a total of 40 database connectors for ICETS-Infosys, utilizing both SQL and NoSQL databases with Python, contributing to the development of valuable internal intellectual property for ICETS(Infosys Center for Emerging Technology Solutions). Created a user-friendly interface for the CFIN tool using Python, which streamlined communication and collaboration with the Infosys SAP Team.
- Managed and maintained 55 repositories within the Enterprise Content Management software Documentum across Production, Integration, and Test environments; automated DAR/JAR deployments, enhancing deployment efficiency by a factor of 16.

Infosys RISE - Insta Award for Excellence
Publications 📚
-
Abstract: Spinal microglia play a pivotal role in the development of neuropathic pain. Peripheral nerve injury induces changes in the transcriptional profile of microglia, including increased expression of components of the translational machinery. Whether microglial protein synthesis is stimulated following nerve injury and has a functional role in mediating pain hypersensitivity is unknown. Here, we show that nascent protein synthesis is upregulated in spinal microglia following peripheral nerve injury in both male and female mice. Stimulating mRNA translation in microglia by selectively ablating the translational repressor eukaryotic initiation factor 4E–binding protein 1 (4E-BP1) promoted the transition of microglia to a reactive state and induced mechanical hypersensitivity in both sexes, whereas spontaneous pain was increased only in males. Conversely, inhibiting microglial translation by expressing a mutant form of 4E-BP1 in microglia attenuated their activation following peripheral nerve injury and alleviated neuropathic pain in both sexes. Thus, stimulating 4E-BP1–dependent translation promotes microglial reactivity and mechanical hypersensitivity, whereas inhibiting it alleviates neuropathic pain.
Show Publication -
Abstract: RNA sequencing studies on human dorsal root ganglion (hDRG) from patients suffering from neuropathic pain show upregulation of OSM, linking this IL-6 family cytokine to pain disorders. In mice, however, OSM signaling causes itch behaviors through a direct effect on its cognate receptor expressed uniquely by pruriceptive sensory neurons. We hypothesized that an expansion in function of OSM-OSM receptor (OSMR) in sensory disorders in humans could be explained by species differences in receptor expression and signaling. Our in situ hybridization and immunohistochemical findings demonstrate broad expression of OSMR in DRG nociceptors and afferent fibers innervating the superficial and deep skin of humans. In patch-clamp electrophysiology, OSM directly activates human sensory neurons engaging MAPK signaling to promote action potential firing. Using CRISPR editing we show that OSM activation of MAPK signaling is dependent on OSMR and not LIFR in hDRG. Bulk, single-nuclei, and single-cell RNA-seq of OSM-treated hDRG cultures reveal expansive similarities in the transcriptomic signature observed in pain DRGs from neuropathic patients, indicating that OSM alone can orchestrate transcriptomic signatures associated with pain. We conclude that OSM-OSMR signaling via MAPKs is a critical signaling factor for DRG plasticity that may underlie neuropathic pain in patients.
Show Publication -
Abstract: Neurons in the dorsal root ganglion (DRG) receive and transmit sensory information from the tissues they innervate and from the external environment. Upper cervical (C1-C2) DRGs are functionally unique as they receive input from the neck, head, and occipital cranial dura, the latter two of which are also innervated by the trigeminal ganglion (TG). The C2 DRG also plays an important role in neck pain, a common and disabling disorder that is poorly understood. Advanced transcriptomic approaches have significantly improved our ability to characterize RNA expression patterns at single-cell resolution in the DRG and TG, but no previous studies have characterized the C2 DRG. Our aim was to use single-nucleus and spatial transcriptomic approaches to create a molecular map of C2 DRGs from patients undergoing arthrodesis surgery with ganglionectomy. Patients with acute (~3 months) or chronic (≥3 months) neck pain were enrolled and completed patient-reported outcomes and quantitative sensory testing prior to surgery. C2 DRGs were characterized with bulk, single nucleus, and spatial RNA sequencing technologies from 22 patients. Through a comparative analysis to published datasets of the lumbar DRG and TG, neuronal clusters identified in both TG and DRG were identified in the C2 DRG. Therefore, our study definitively characterizes the molecular composition of human C2 neurons and establishes their similarity with unique characteristics of subsets of TG neurons. We identified differentially expressed genes in endothelial, fibroblast and myelinating Schwann cells associated with chronic pain, including FGFBP2, C8orf34 and EFNA1 which have been identified in previous genome and transcriptome wide association studies (GWAS/TWAS). Our work establishes an atlas of the human C2 DRG and identifies altered gene expression patterns associated with chronic neck pain. This work establishes a foundation for the exploration of painful disorders in humans affecting the cervical spine.
Show Publication -
Abstract: Human dermal sleeping nociceptors display ongoing activity in neuropathic pain, affecting 10% of the population. Despite advances in rodents, a molecular marker for these mechano-insensitive C-fibers (CMis) in human skin remains elusive, preventing targeted therapy. In this translational Patch-seq study, we combine single-cell transcriptomics following electrophysiological characterization with single-nucleus and spatial transcriptomics from pigs and humans. We functionally identified CMis in pig sensory neurons with patch-clamp using adapted protocols from human microneurography. We identified oncostatin-M-receptor (OSMR) and somatostatin (SST) as marker genes for CMis. Following dermal injection in healthy human volunteers, oncostatin-M, the ligand of OSMR, exclusively modulates CMis. We identified the entire molecular architecture of human dermal sleeping nociceptors, providing new therapeutic targets and the basis for a mechanistic understanding of neuropathic pain.
Show Publication -
Abstract: Sensitization of spinal nociceptive circuits plays a crucial role in neuropathic pain. This sensitization depends on new gene expression that is primarily regulated via transcriptional and translational control mechanisms. The relative roles of these mechanisms in regulating gene expression in the clinically relevant chronic phase of neuropathic pain are not well understood. Here, we show that changes in gene expression in the spinal cord during the chronic phase of neuropathic pain are substantially regulated at the translational level. Downregulating spinal translation at the chronic phase alleviated pain hypersensitivity. Cell-type-specific profiling revealed that spinal inhibitory neurons exhibited greater changes in translation after peripheral nerve injury compared to excitatory neurons. Notably, increasing translation selectively in all inhibitory neurons or parvalbumin-positive (PV+) interneurons, but not excitatory neurons, promoted mechanical pain hypersensitivity. Furthermore, increasing translation in PV+ neurons decreased their intrinsic excitability and spiking activity, whereas reducing translation in spinal PV+ neurons prevented the nerve injury-induced decrease in excitability. Thus, translational control mechanisms in the spinal cord, particularly in inhibitory neurons, play a role in mediating neuropathic pain hypersensitivity.
Show Publication -
Abstract: Diabetic peripheral neuropathy (DPN) is a prevalent complication of diabetes mellitus that is caused by metabolic toxicity to peripheral axons. We aimed to gain deep mechanistic insight into the disease process using bulk and spatial RNA sequencing on tibial and sural nerves recovered from lower leg amputations in a mostly diabetic population. First, our approach comparing mixed sensory and motor tibial and purely sensory sural nerves shows key pathway differences in affected nerves, with distinct immunological features observed in sural nerves. Second, spatial transcriptomics analysis of sural nerves reveals substantial shifts in endothelial and immune cell types associated with severe axonal loss. We also find clear evidence of neuronal gene transcript changes, like PRPH, in nerves with axonal loss suggesting perturbed RNA transport into distal sensory axons. This motivated further investigation into neuronal mRNA localization in peripheral nerve axons generating clear evidence of robust localization of mRNAs such as SCN9A and TRPV1 in human sensory axons. Our work gives new insight into the altered cellular and transcriptomic profiles in human nerves in DPN and highlights the importance of sensory axon mRNA transport as an unappreciated potential contributor to peripheral nerve degeneration.
Show Publication -
Abstract: Gene expression is influenced by chromatin architecture via controlled access of regulatory factors to DNA. To better understand gene regulation in the human dorsal root ganglion (hDRG) we used bulk and spatial transposase-accessible chromatin technology followed by sequencing (ATAC-seq). Using bulk ATAC-seq, we detected that in females diverse differentially accessible chromatin regions (DARs) mapped to the X chromosome and in males to autosomal genes. EGR1/3 and SP1/4 transcription factor binding motifs were abundant within DARs in females, and JUN, FOS and other AP-1 factors in males. To dissect the open chromatin profile in hDRG neurons, we used spatial ATAC-seq. The neuron cluster showed higher chromatin accessibility in GABAergic, glutamatergic, and interferon-related genes in females, and in Ca2+- signaling-related genes in males. Sex differences in transcription factor binding sites in neuron-proximal barcodes were consistent with the trends observed in bulk ATAC-seq data. We validated that EGR1 expression is biased to female hDRG compared to male. Strikingly, XIST, the long-noncoding RNA responsible for X inactivation, hybridization signal was found to be highly dispersed in the female neuronal but not non-neuronal nuclei suggesting weak X inactivation in female hDRG neurons. Our findings point to baseline epigenomic sex differences in the hDRG that likely underlie divergent transcriptional responses that determine mechanistic sex differences in pain.
Show Publication -
Abstract: Diabetic Peripheral Neuropathy (DPN) stands out as a very common and clinically consequential complication of diabetes. DPN is characterized by nerve conduction impairment, dieback of nerve endings from the skin, spontaneous pain, and numbness in extremities. In this study, we conducted single-nucleus RNA sequencing on human dorsal root ganglion (DRG) tissues recovered from organ donors with a documented history of diabetes and neuropathic pain. Employing 10x Genomics technology, we performed single-nucleus RNA sequencing on eight human DRG samples each from individuals with DPN or healthy donors. Our primary objectives encompassed elucidating distinct populations of neurons and comprehending their transcriptomic variations between healthy and DPN DRGs. Additionally, we sought to characterize the diversity of transcriptomic changes in non-neuronal cells and explore their neuro-immune interactions in DPN. Our analysis revealed significant alterations in non-neuronal cell populations, including T cells, macrophages, and adipocytes within human DRG of DPN donors. Our work presents a comprehensive single-cell transcriptomic landscape of human DRGs in DPN donors, providing insights into molecular profile changes associated with DPN. The interactions between neuronal and non-neuronal cells could offer valuable insights into the underlying mechanisms of this condition, assisting in the development of novel therapeutics to alleviate DPN. Funded by National Institute of Health (U19 NS130608).
Show Publication -
Abstract: Hyperalgesic priming is a model system that has been widely used to understand plasticity in painful stimulus-detecting sensory neurons, called nociceptors. A key feature of this model system is that following priming, stimuli that do not normally cause hyperalgesia now readily provoke this state. We hypothesized that hyperalgesic priming occurs due to reorganization of translation of mRNA in nociceptors. To test this hypothesis, we used paclitaxel treatment as the priming stimulus and translating ribosome affinity purification (TRAP) to measure persistent changes in mRNA translation in Nav1.8+ nociceptors. TRAP sequencing revealed 161 genes with persistently altered mRNA translation in the primed state. We identified Gpr88 as upregulated and Metrn as downregulated. We confirmed a functional role for these genes, wherein a GPR88 agonist causes pain only in primed mice and established hyperalgesic priming is reversed by Meteorin. Our work demonstrates that altered nociceptor translatomes are causative in producing hyperalgesic priming.
Show Publication The Journal of the International Association for the Study of Pain Download Paper
Skills Journey ⚒️
Generative AI
Langchain, Langgraph, crewAI, LangLarge Language Models (LLMs), Custom fine-tuning LLMs, Evals(DeepEval, LangSmith) Prompt Engineering(Meta prompting and prompt folding), Retrieval Augmented Generation(RAG), Agentic-RAG, HuggingFace smolagents, LLMOps, Attention Mechanisms, Diffusion Models, GANs (expand for more)
Experience building and deploying generative AI solutions for text, voice and vision, image generation, including custom RAG pipelines and multi-agent systems.
Machine Learning and AI
Data Science, Machine Learning, Computer Vision and Deep Learning, Natural Language Processing: RNNs, LSTMs, BERT and BART, Transformers, Computer Vision models, Tensorflow, PyTorch, Keras, Folium, Plotly & Bokeh Visuvalization Tools
Experienced in statistical analysis, data visualization, and predictive modeling.
Programming
Python, R-programming, Shell Scripting, Java, SQL, NoSQL, DQL, Javascript, HTML, CSS
Proficient in developing scalable applications, application endpoints, load-balancing(Nginx) and automating workflows.
Tools & Technologies
Cloud Computing: Azure(Certified) and AWS(Certified), Docker, Kubernetes, Cromwell, Nextflow, Nginx, CI/CD Pipelines, Documentum (ECM Software), Agile Certified Practitioner
Experienced in versioning of ML models, tackling model and data drifts, deployments, containerization, orchestration, and workflow management.
Web Servers
Tornado, Django, Flask
Skilled in building and deploying web applications and APIs.
Other Skills
Bioinformatics, Omics, Genomics, Transcriptomics, Bulk RNA-sequemcing Analysis (DE Genes), Visium Analysis(Spatial Transcriptomics Data), ATAC-sequencing, Single-cell Analysis, AAV Capsid Engineering
Experienced in Bioinformatics and Neuroinformatics Analysis.
Projects 📁

AI Powered Travel Agent using LangChain
An AI‑powered travel planner that auto‑generates day‑by‑day itineraries, budget breakdowns, and live weather forecasts—accessible via both a CLI and an interactive Streamlit web app.
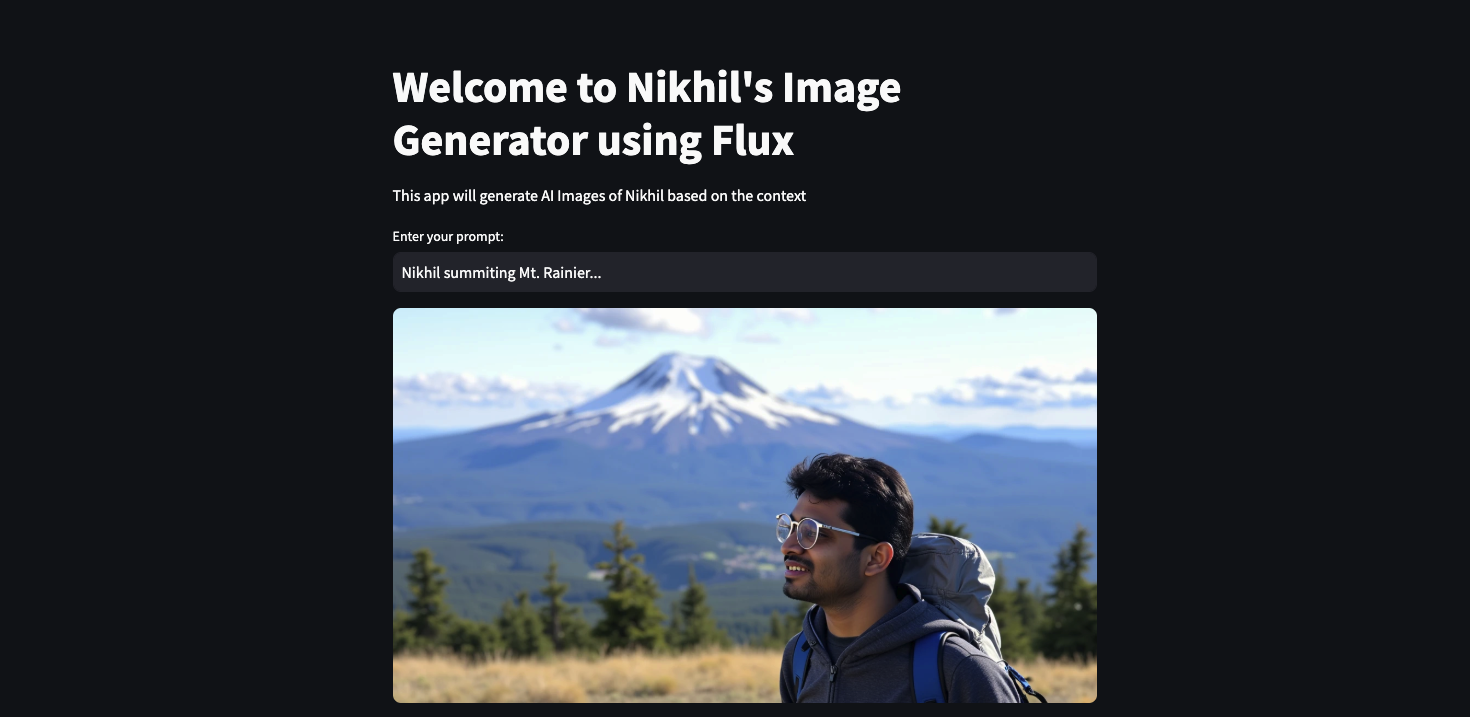
Image Generator using FLUX
FLUX-image-generator is an AI-powered image generation tool that utilizes the Flux framework and Gemini API to create personalized images from user prompts.

NikhilRAG
NikhilRAG is an AI-powered RAG Chatbot that dynamically synthesizes my professional journey, projects, and skills. Powered by GPT-LangChain and Streamlit.
Xenium Cell Segmentation Pipeline
End-to-end pipeline for spinal-cord microscopy segmentation with Cellpose, tiling, and mask stitching.

Agentic (RAG) Retrieval Augmented Generation
This isn't just another RAG implementation—it's an agentic system that intelligently decides when and how to retrieve information.

S.A.G.E - Strategic Analysis & Guidance Engine
SAGE is an AI-powered strategist, researcher, and builder all rolled into one. It is a swarm of 13 crews, decoupled for processing Human-in-the-loop requests and versioning runs.
View on GitHub →
Smart ATS Resume crewAI multi-agent system
An AI-powered resume tailoring system that automatically optimizes your resume for specific job posting.
View on GitHub →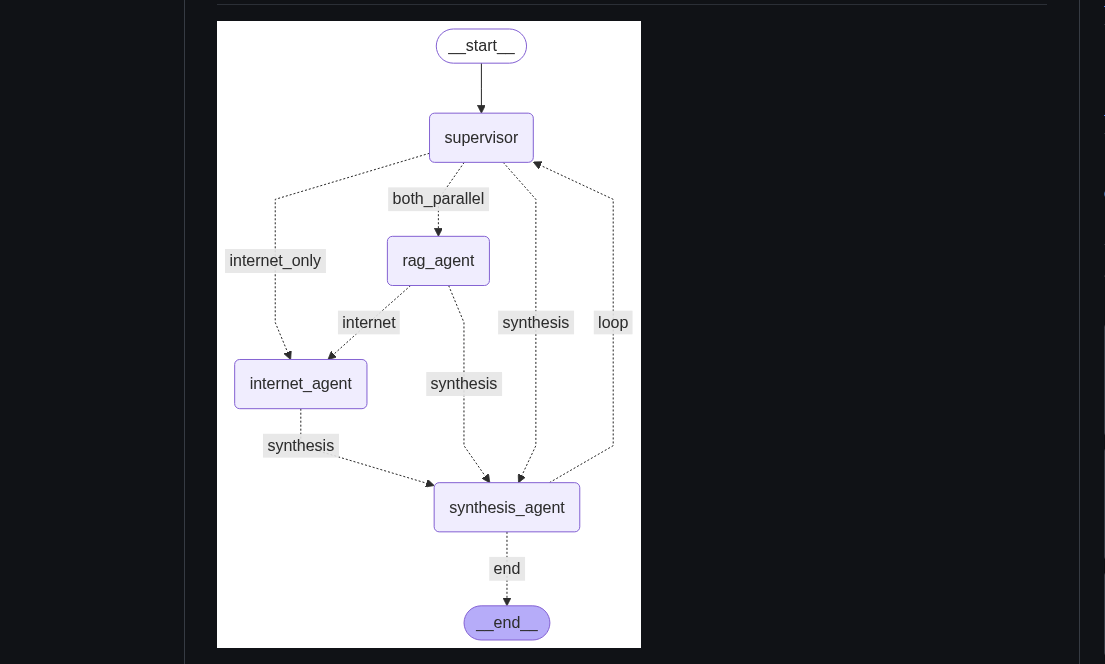
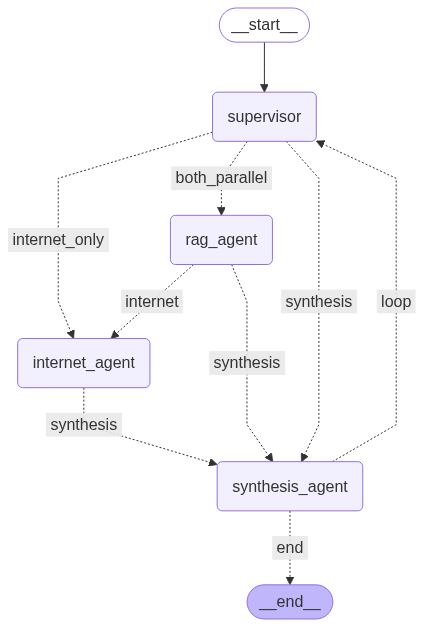
Multi-agent Research RAG
A multi-agent chatbot combining Retrieval-Augmented Generation (RAG) with live internet search for concise, up-to-date answers.
View on GitHub →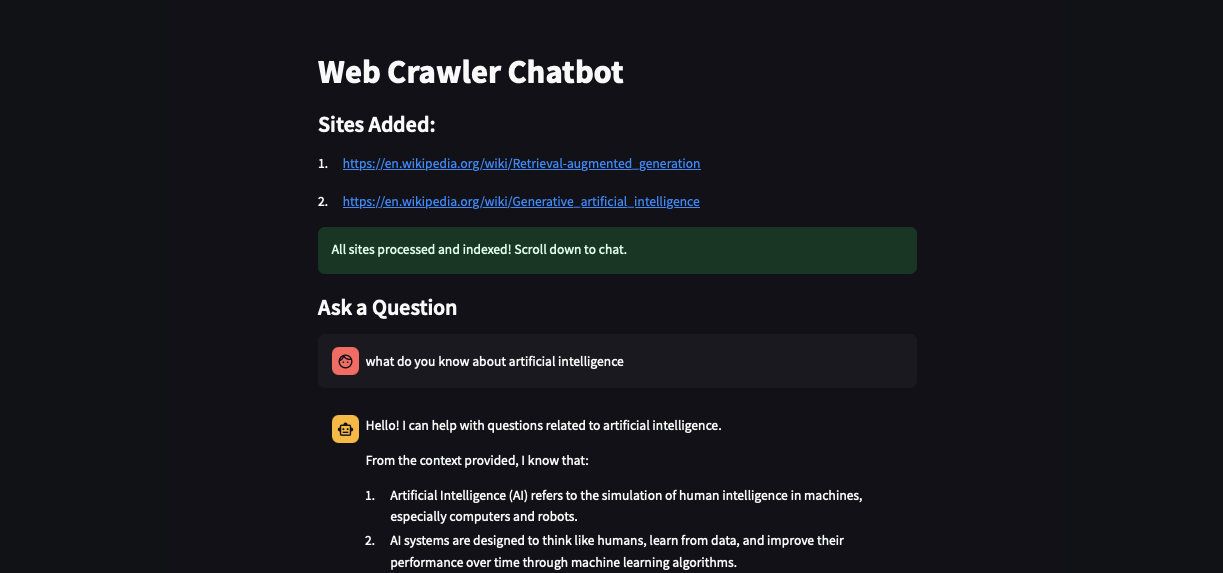
RAG Web-Scraper Chatbot using Ollama(local)
A powerful web crawler chatbot that extracts, indexes, and intelligently answers questions on web content using Selenium, LangChain, and Ollama's LLM.
View on GitHub →
Spotify Wrapped Unwrapped
Spotify Unwrapped — A Streamlit web app that connects to your Spotify account, visualizes your top tracks & artists, and delivers GPT-powered lyrical, emotional, and psychological insights into your listening habits.
View on GitHub →
YouTube CrewAI Scraper & Mapper (Agentic scraper for YouTube videos)
A Python tool that uses crewAI to scrape YouTube channel data and visualizes it geographically with interactive maps.
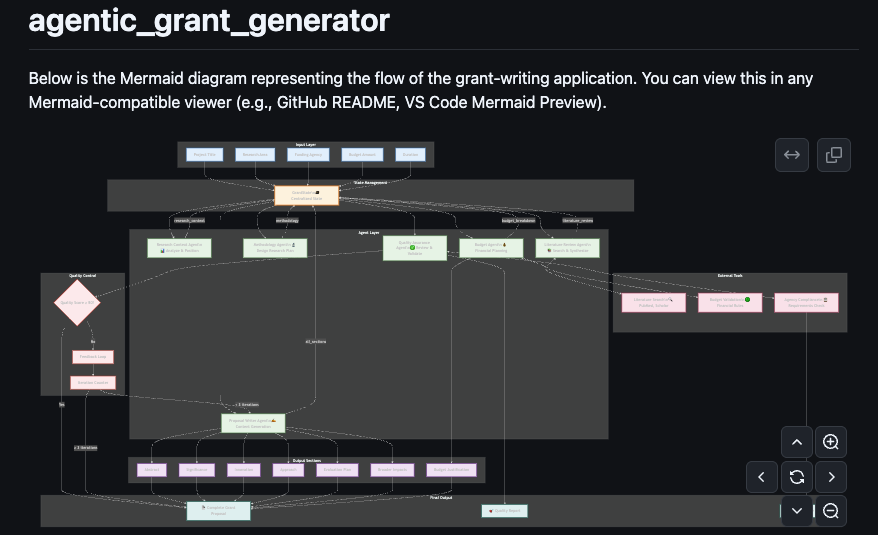
Agentic Grant Generator
Smart Agentic Grant Generator that pilots a draft version of grants.
View on GitHub →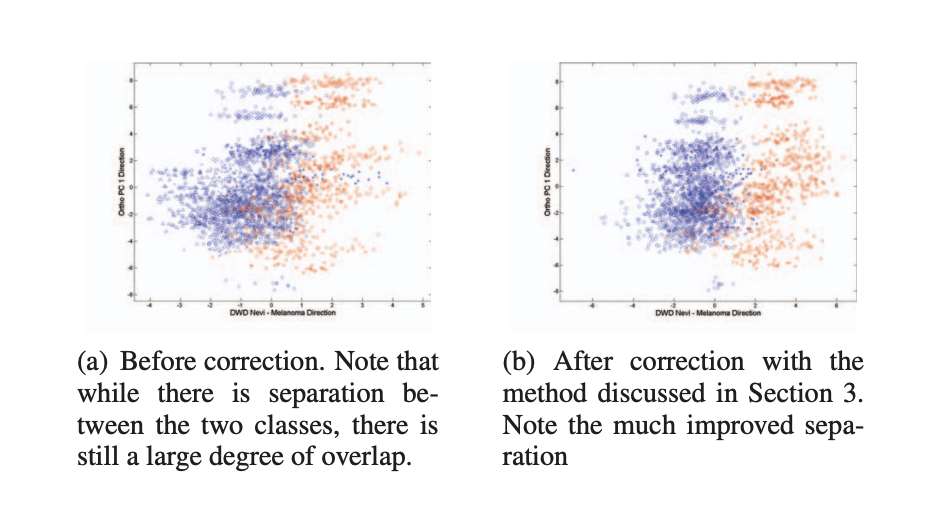
Normalize H&E images for 10X-Visium workflows
A method to normalize H and E images for Image Processing and Quantitative Testing, this is referenced from a paper.
View on GitHub →
TRAP Analysis Tool
TRAP-Analysis tool enables the investigation of cell-specific mRNA associations with ribosomes obtained from Translating Ribosome Affinity Purification (TRAP) sequencing.
View on GitHub →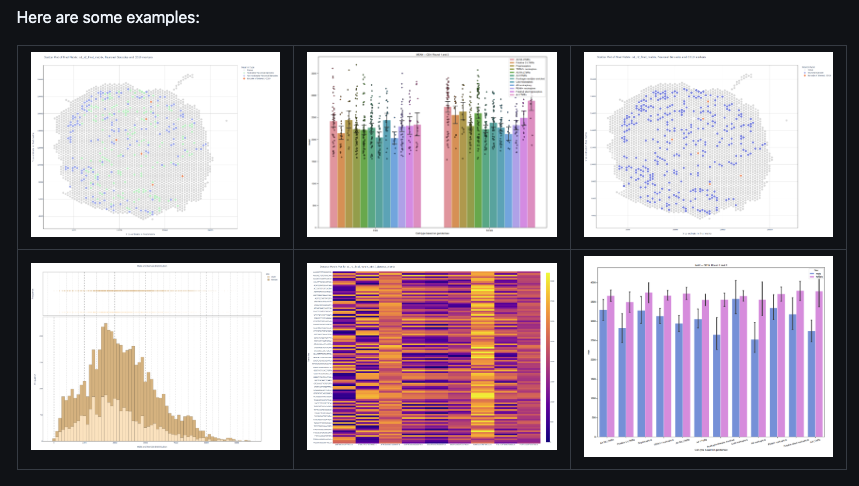
Visium Distance Plotting Tool (Euclidian Distances)
The Visium Distance Plotting Tool is a python-based application that offers a range of interactive plots for analyzing visium single-cell data(e.g. csv files).
View on GitHub →
Visium Spatial Analysis
Visium Spatial Gene Expression is a next-generation molecular profiling solution for classifying tissue based on total mRNA. The steps involved in classifying the Spatial Gene Expression is done in this tool using python and R.
View on GitHub →
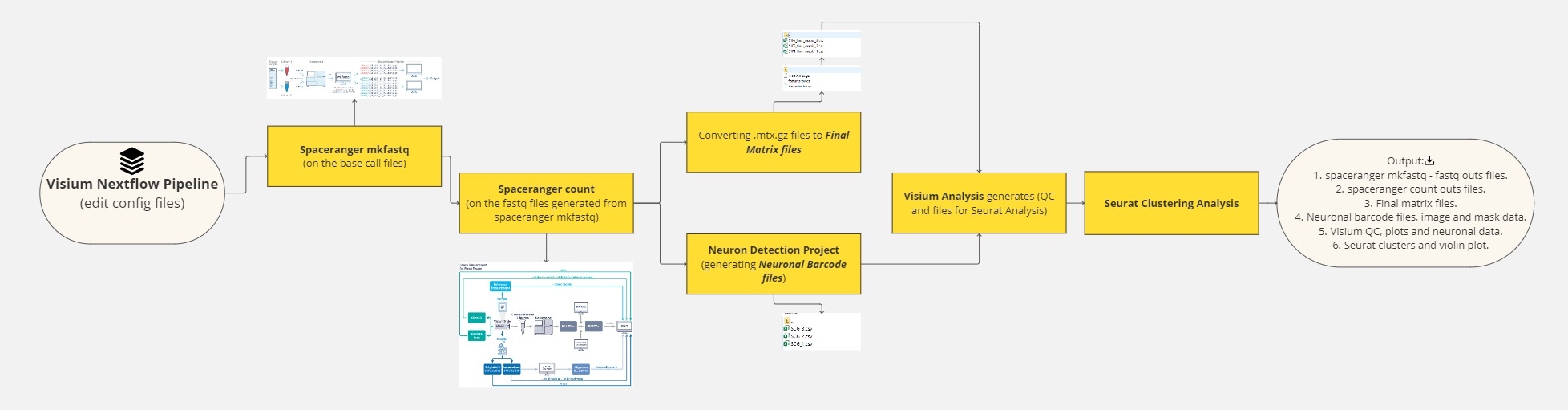
Visium Spatial Analysis using NextFlow
This is an automated workflow pipeline for analyzing Visium data, implemented in Python, R, and Shell scripts, and wrapped in a NextFlow workflow.
View on GitHub →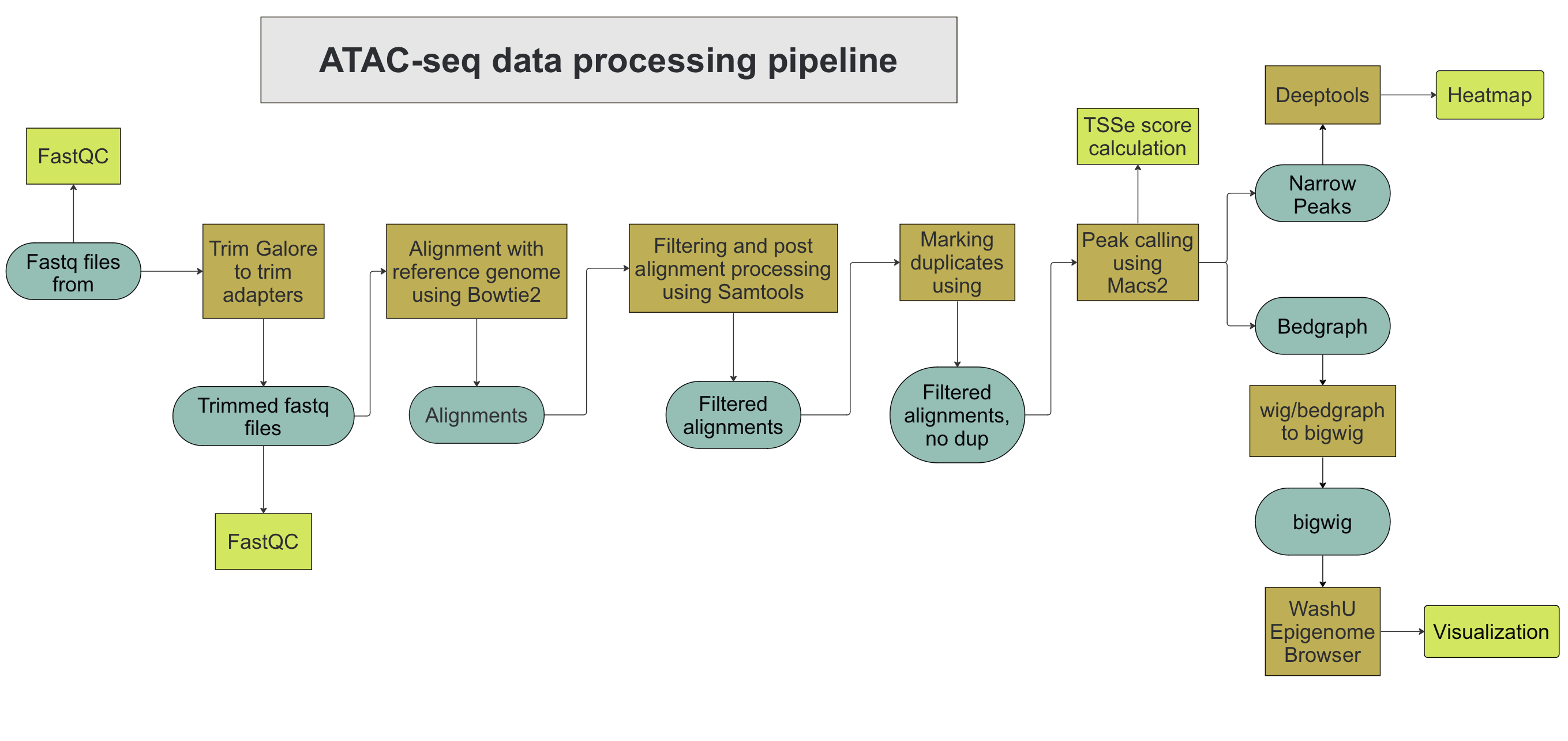
ATACSeq Data Processing Pipeline using NextFlow
This is an automated workflow pipeline for analyzing bulk ATAC-seq data, implemented primarily in bash scripts, and wrapped in a NextFlow workflow.
View on GitHub →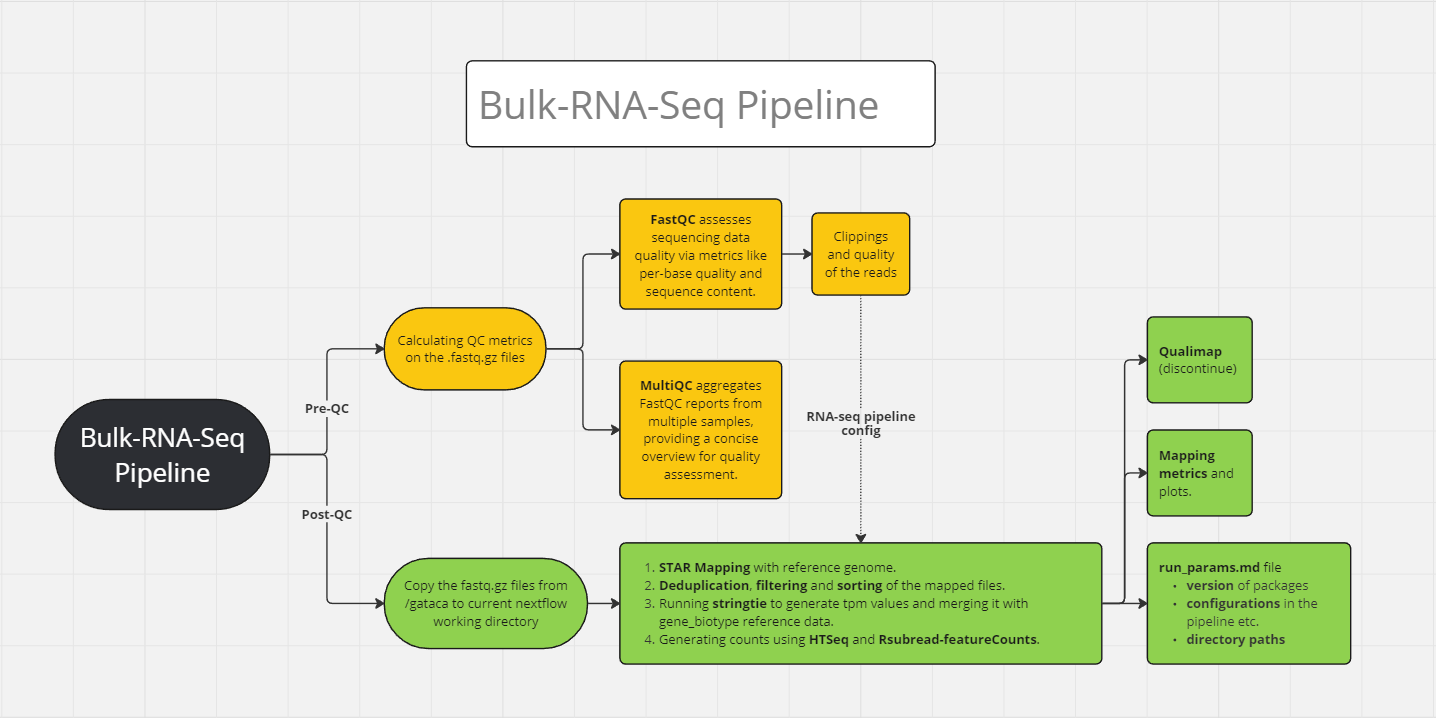
Bulk-RNA Seq Data Processing Pipeline using NextFlow / Docker
This is an automated workflow pipeline for analyzing and processing Bulk-RNA seq data, implemented primarily in bash, python and R, and wrapped in a NextFlow workflow to characterize the gene landscape in the samples.
View on GitHub →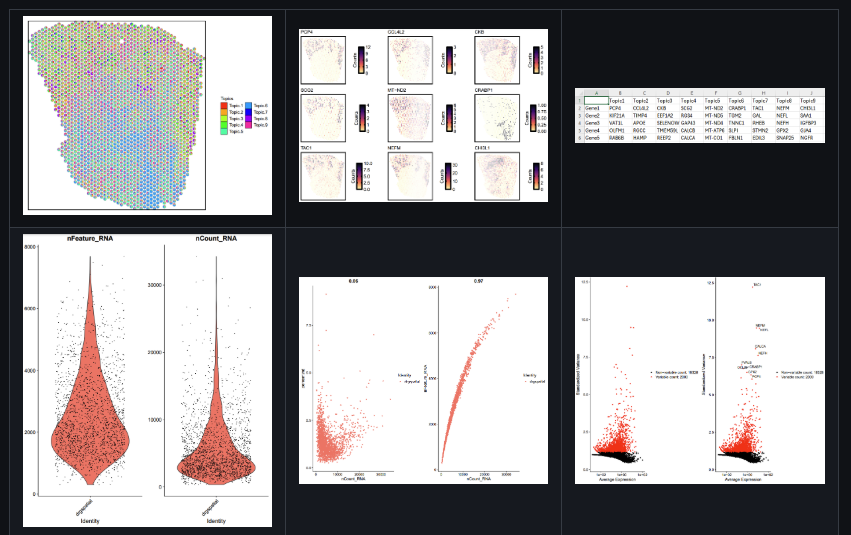
Analysis Pipeline for Visium 10x Data [visium10x-spatial-transcriptomics-pipeline]
This repository provides an R-based pipeline for analyzing spatial RNA-seq data using Seurat, STDeconvolve, and Giotto. Designed to process and analyze spatial transcriptomics data through a sequence of well-defined steps, enabling in-depth exploration and interpretation of spatially resolved gene expression profiles.
View on GitHub →
Xenium Cell Segmentation using YOLO-v11
Cell segmentation - xenium data: human spinal cord, hDRG samples.
View on GitHub →Education 🎓
The University of Texas at Dallas
Master's in Business Analytics and Artificial Intelligence
Jan 2023 - Present
Ramaiah Institute of Technology
Bachelor of Engineering in Mechanical Engineering
Aug 2014 - June 2018
Certifications 📜

May 2024
![Introduction to DevOps [IBM]](images/ibm_devops.png)
Jul 2024
Articles 📚

Beyond the Words: How Prompt Engineering … Builds Exceptional AI Agents
Crafting prompts, agentic workflows, and evaluation loops that turn LLMs from basic responders into powerful research assistants.
Read on Medium →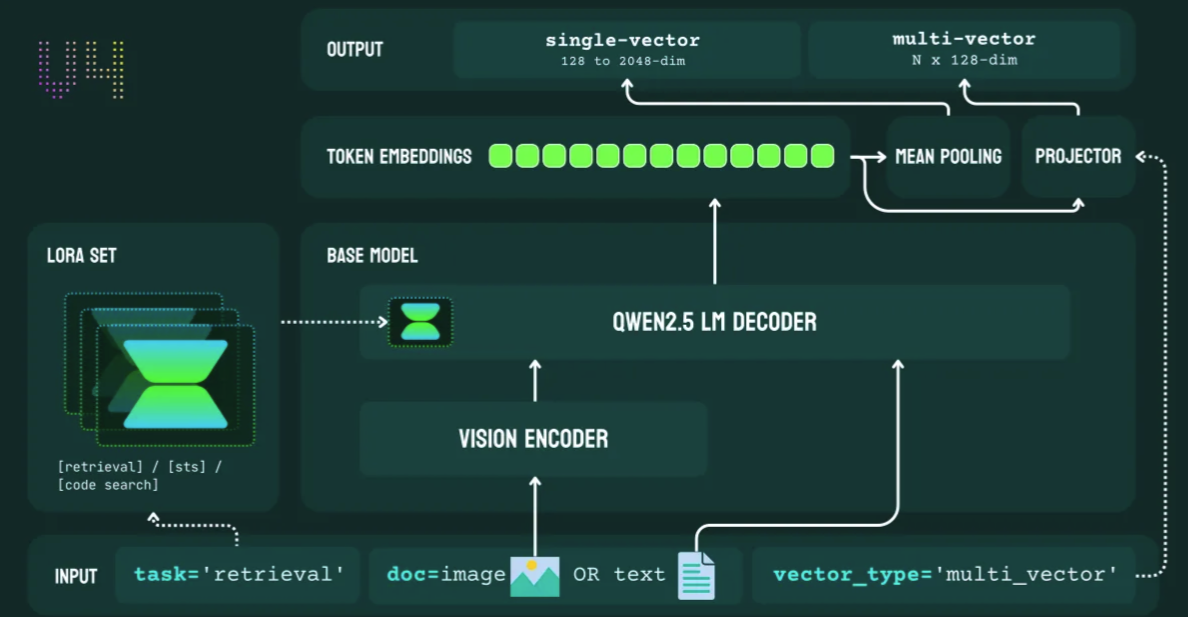
Jina Embeddings v4: One Model to Embed Them All
3.8-B param multimodal backbone, 32 K tokens, Matryoshka vectors & state-of-the-art retrieval in 30 languages.
Read on Medium →